Recipes¶
Worked examples using the grammar API. Each recipe starts from data and builds up to a finished chart. The progression runs from simple single-mark charts to multi-layer compositions with per-group transforms.
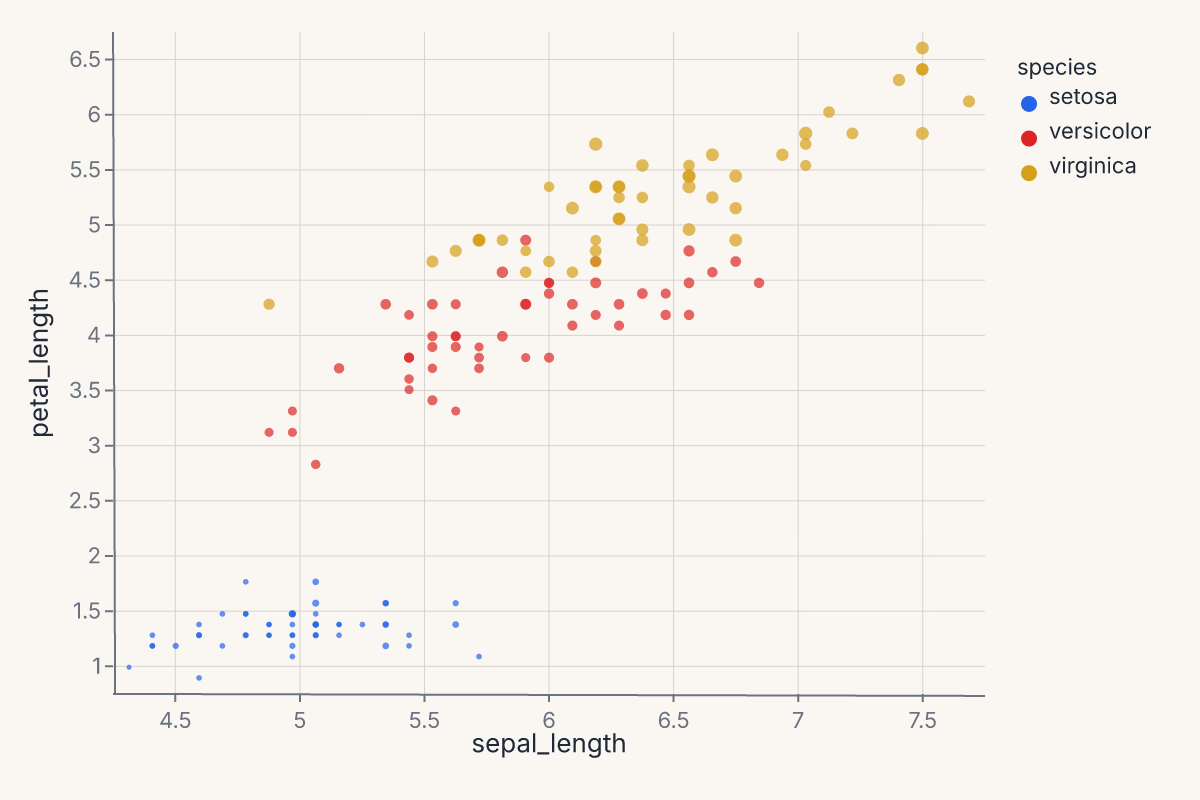
Scatter with color and size¶
Map two continuous fields to position and two more to appearance:
import ferrum as fm
import polars as pl
from sklearn.datasets import load_iris
raw = load_iris()
iris = pl.DataFrame(raw.data, schema=["sepal_length", "sepal_width", "petal_length", "petal_width"]).with_columns(
species=pl.Series([raw.target_names[t] for t in raw.target])
)
chart = (
fm.Chart(iris)
.mark_point(opacity=0.7)
.encode(
x="sepal_length",
y="petal_length",
color="species:N",
size="petal_width",
)
)
assert chart.show_svg().startswith("<svg")
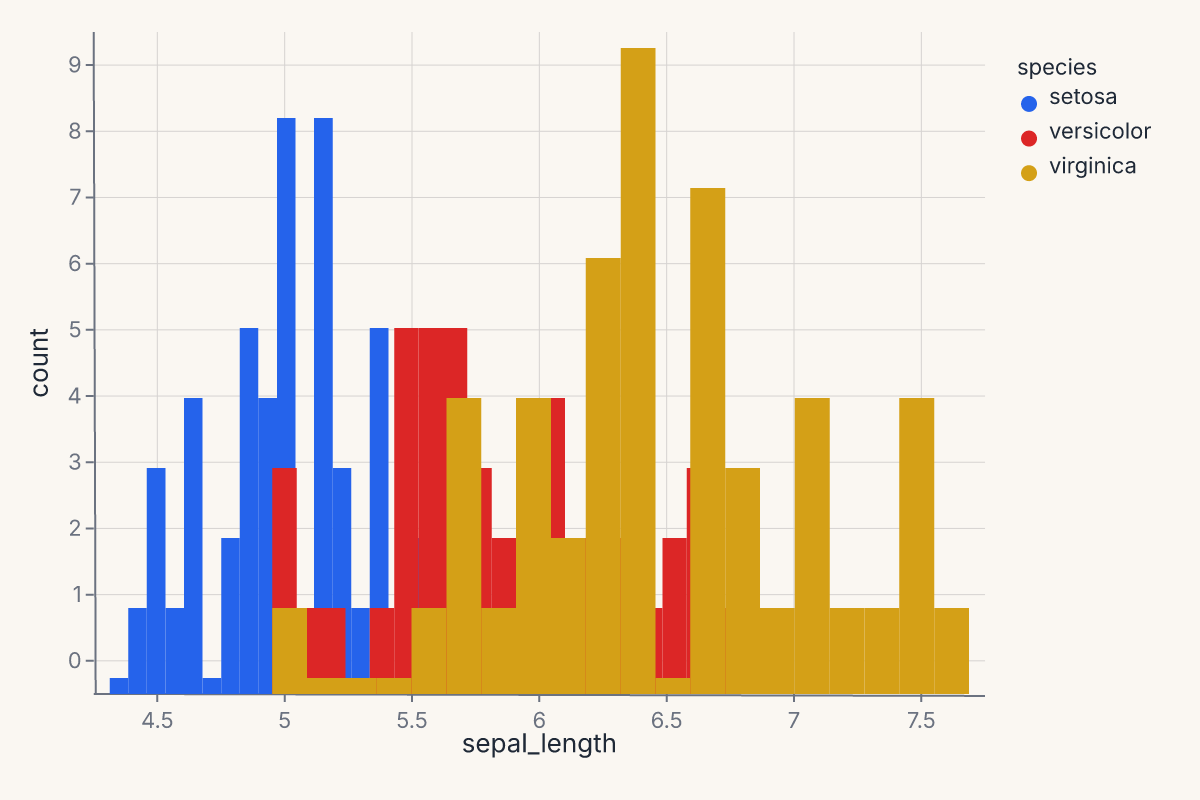
Histogram with hue overlay¶
Stack or layer distributions by a categorical variable:
import ferrum as fm
import polars as pl
from sklearn.datasets import load_iris
raw = load_iris()
iris = pl.DataFrame(raw.data, schema=["sepal_length", "sepal_width", "petal_length", "petal_width"]).with_columns(
species=pl.Series([raw.target_names[t] for t in raw.target])
)
chart = (
fm.Chart(iris)
.mark_histogram(bin_count=20, groupby="species")
.encode(x="sepal_length", color="species:N")
)
assert chart.show_svg().startswith("<svg")
Per-group KDE¶
Use groupby to compute a separate density estimate per category:
import ferrum as fm
import polars as pl
from sklearn.datasets import load_iris
raw = load_iris()
iris = pl.DataFrame(raw.data, schema=["sepal_length", "sepal_width", "petal_length", "petal_width"]).with_columns(
species=pl.Series([raw.target_names[t] for t in raw.target])
)
chart = (
fm.Chart(iris)
.mark_density(bandwidth="scott", groupby="species")
.encode(x="sepal_length", color="species:N")
)
assert chart.show_svg().startswith("<svg")
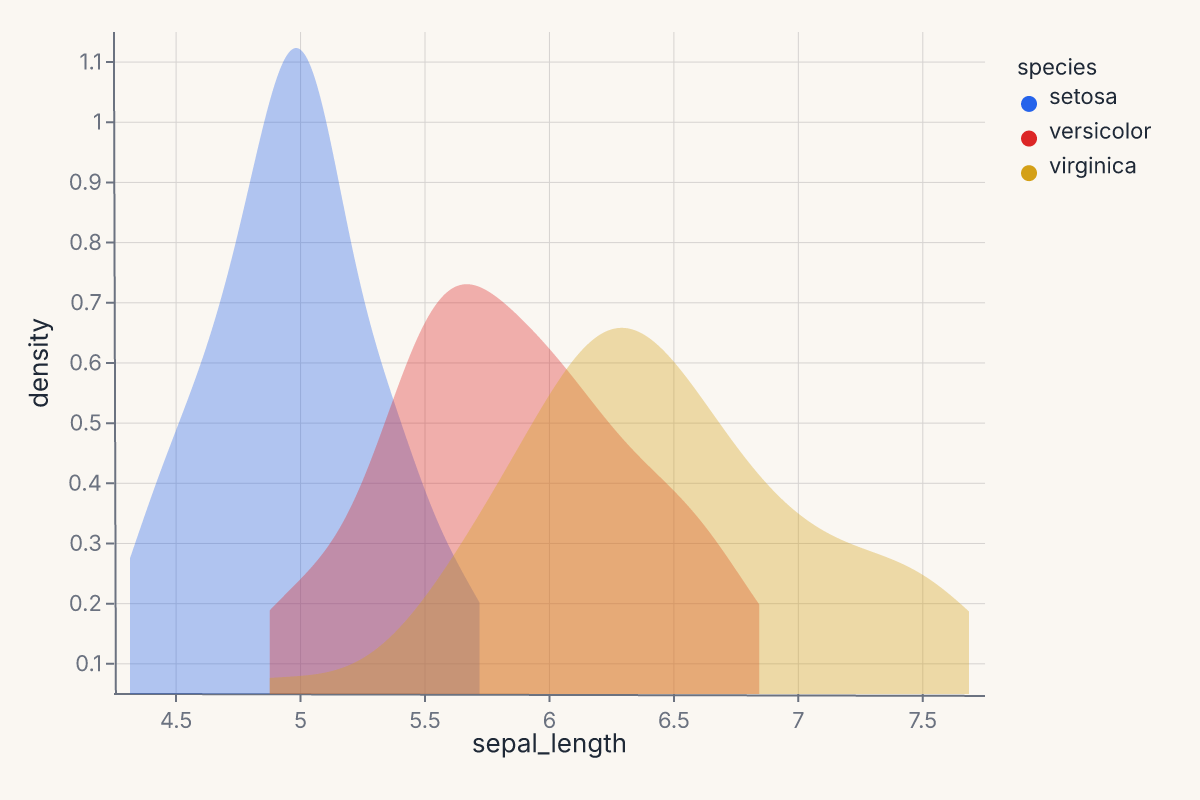
Without groupby, the density is computed over all rows combined. With groupby="species", each species gets its own KDE curve colored by the color encoding.
Scatter + per-group smooth¶
Layer a scatter with per-species LOESS trend lines. The groupby parameter is essential — without it, one combined line is computed:
import ferrum as fm
import polars as pl
from sklearn.datasets import load_iris
raw = load_iris()
iris = pl.DataFrame(raw.data, schema=["sepal_length", "sepal_width", "petal_length", "petal_width"]).with_columns(
species=pl.Series([raw.target_names[t] for t in raw.target])
)
points = (
fm.Chart(iris)
.mark_point(opacity=0.6)
.encode(x="sepal_length", y="petal_length", color="species:N")
)
trend = (
fm.Chart(iris)
.mark_smooth(method="loess", groupby="species")
.encode(x="sepal_length", y="petal_length", color="species:N")
)
chart = points + trend
assert chart.show_svg().startswith("<svg")
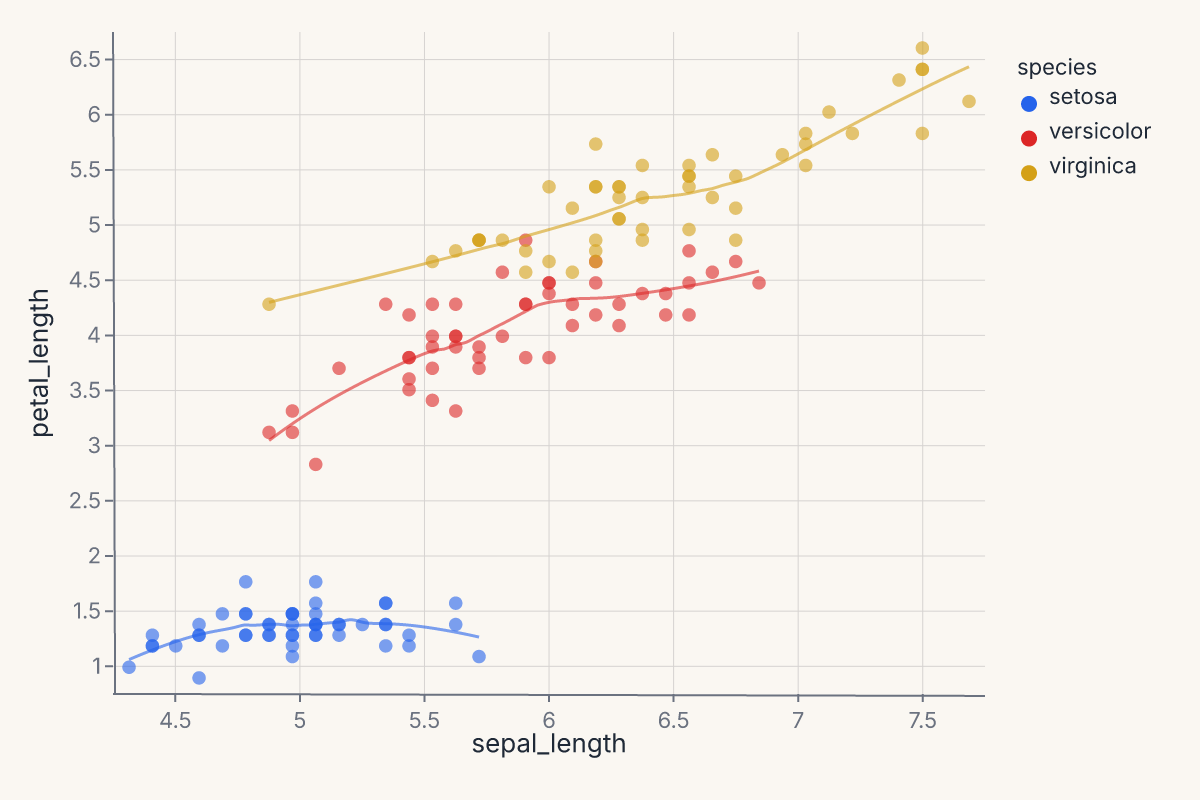
The + operator layers both marks on shared axes. The scatter renders the raw points; the smooth renders the fitted curves. Both share the same color scale.
Scatter + smooth with confidence interval¶
Add a confidence band around the regression line with ci=:
import ferrum as fm
import polars as pl
from sklearn.datasets import load_iris
raw = load_iris()
iris = pl.DataFrame(raw.data, schema=["sepal_length", "sepal_width", "petal_length", "petal_width"]).with_columns(
species=pl.Series([raw.target_names[t] for t in raw.target])
)
points = (
fm.Chart(iris)
.mark_point(opacity=0.5)
.encode(x="sepal_length", y="petal_length")
)
fit = (
fm.Chart(iris)
.mark_smooth(method="lm", ci=0.95)
.encode(x="sepal_length", y="petal_length")
)
chart = points + fit
assert chart.show_svg().startswith("<svg")
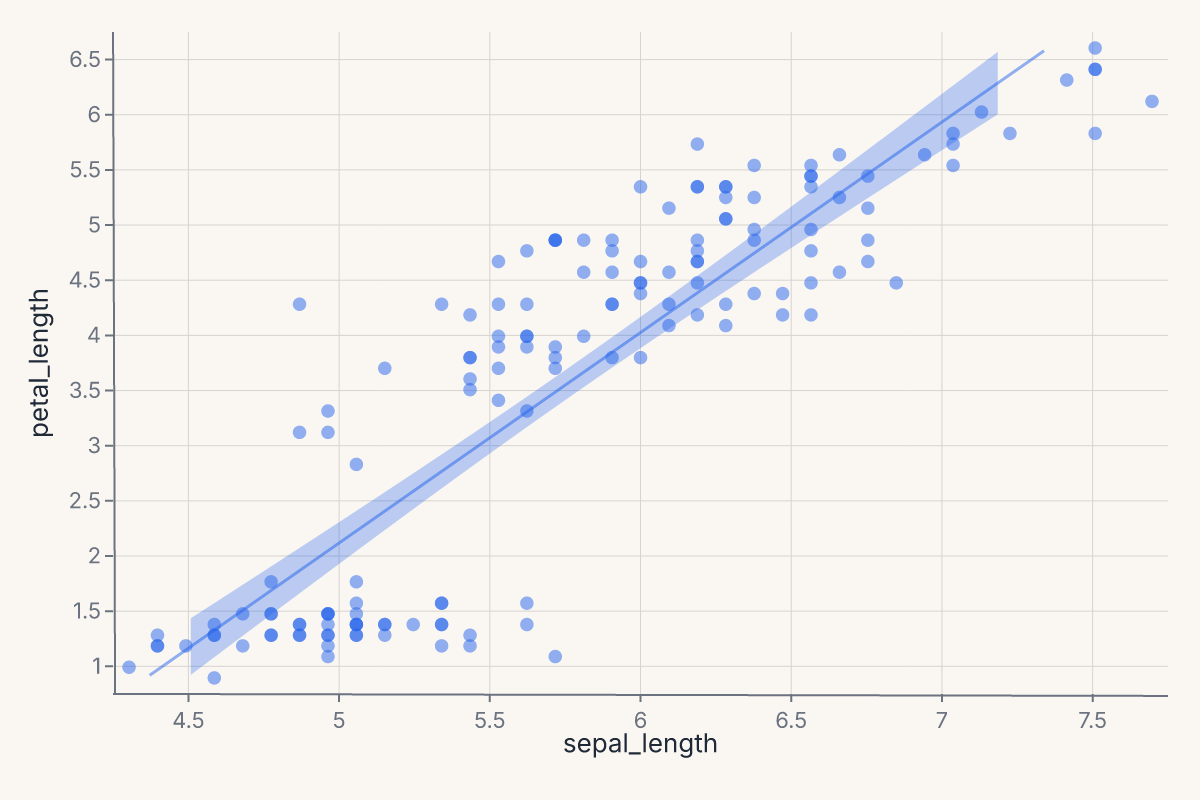
ci=0.95 produces a layered ribbon (band) + line chart. The ribbon shows the 95% confidence interval around the OLS fit. Use method="lm" for linear regression or method="loess" for a local smoother.
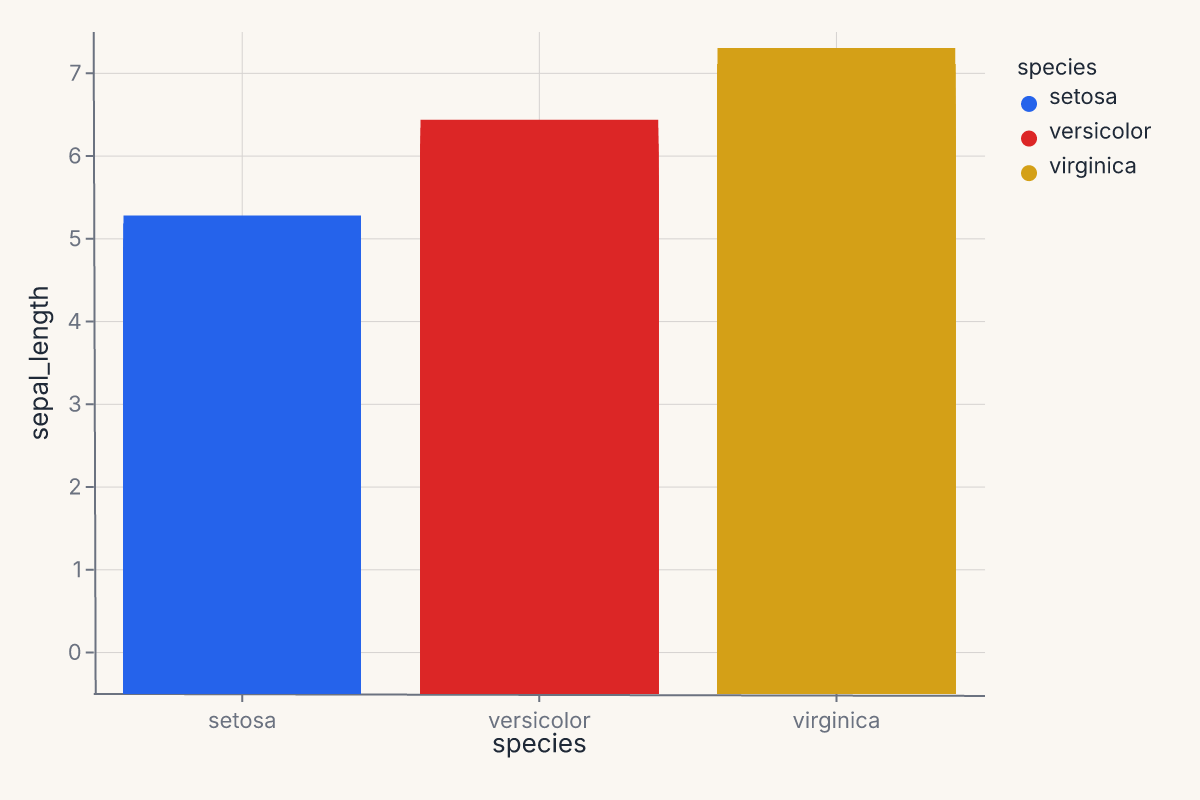
Bar chart with color¶
A simple grouped bar chart using a categorical x-axis:
import ferrum as fm
import polars as pl
from sklearn.datasets import load_iris
raw = load_iris()
iris = pl.DataFrame(raw.data, schema=["sepal_length", "sepal_width", "petal_length", "petal_width"]).with_columns(
species=pl.Series([raw.target_names[t] for t in raw.target])
)
chart = (
fm.Chart(iris)
.mark_bar()
.encode(x="species:N", y="sepal_length", color="species:N")
)
assert chart.show_svg().startswith("<svg")
Pairwise scatter grid¶
Use RepeatChart to show multiple field combinations in a grid:
import ferrum as fm
import polars as pl
from sklearn.datasets import load_iris
raw = load_iris()
iris = pl.DataFrame(raw.data, schema=["sepal_length", "sepal_width", "petal_length", "petal_width"]).with_columns(
species=pl.Series([raw.target_names[t] for t in raw.target])
)
template = (
fm.Chart(iris)
.mark_point(opacity=0.6)
.encode(x=fm.Repeat.column, y=fm.Repeat.row, color="species:N")
)
chart = fm.RepeatChart(
template,
row=["sepal_length", "petal_length"],
column=["sepal_width", "petal_width"],
)
assert chart.show_svg().startswith("<svg")
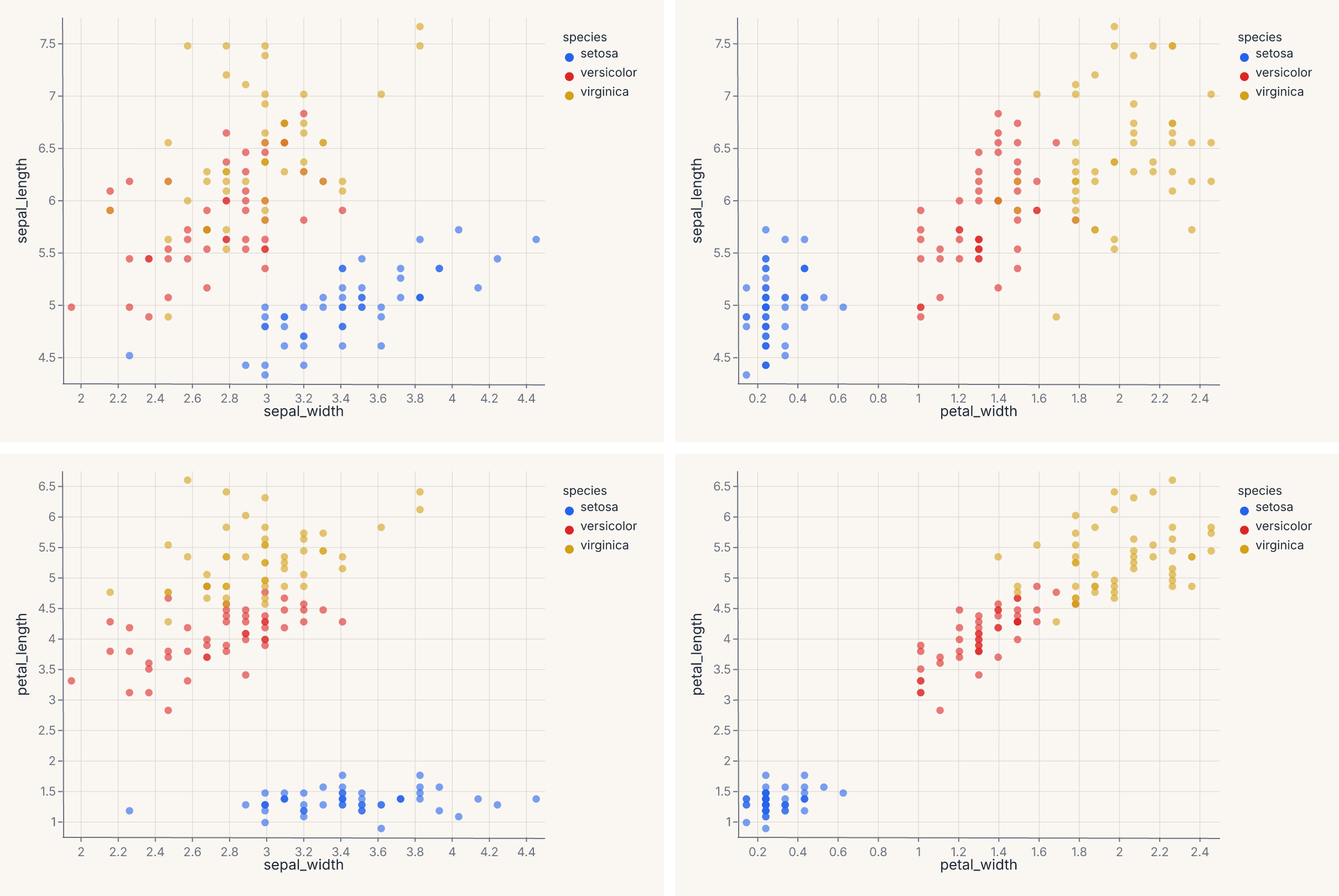
Each cell in the grid swaps one field into the x or y encoding. This is the grammar-level equivalent of fm.pairplot() — use RepeatChart when you want full control over the template chart.
Three-layer scatter with smooth and rug¶
Stack three grammar layers into one chart — scatter, per-group LOESS trend, and a rug plot showing the marginal distribution:
import ferrum as fm
import polars as pl
from sklearn.datasets import load_iris
raw = load_iris()
iris = pl.DataFrame(raw.data, schema=["sepal_length", "sepal_width", "petal_length", "petal_width"]).with_columns(
species=pl.Series([raw.target_names[t] for t in raw.target])
)
points = (
fm.Chart(iris)
.mark_point(opacity=0.5, size=40)
.encode(x="sepal_length", y="petal_length", color="species:N")
)
smooth = (
fm.Chart(iris)
.mark_smooth(method="loess", groupby="species")
.encode(x="sepal_length", y="petal_length", color="species:N")
)
rug = (
fm.Chart(iris)
.mark_tick(opacity=0.3)
.encode(x="sepal_length", color="species:N")
)
chart = points + smooth + rug
assert chart.show_svg().startswith("<svg")
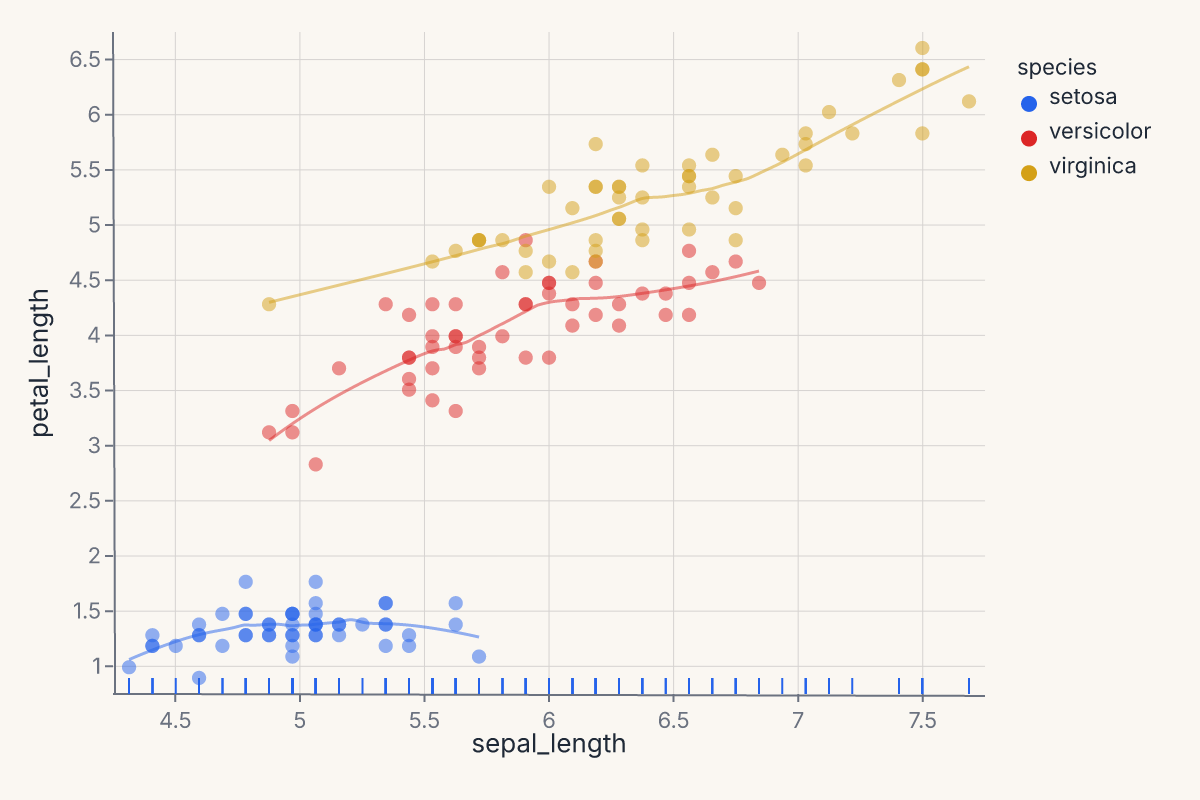
Each + adds another layer on the same axes. All three layers share the x/y scales and the color palette. The rug ticks along the bottom show the marginal distribution of sepal length per species.
Multi-panel model report¶
Compose multiple diagnostic charts into a single view. Train a model, then build a four-panel report with one line of composition:
import ferrum as fm
import polars as pl
import numpy as np
from sklearn.datasets import load_iris
from sklearn.ensemble import RandomForestClassifier
from sklearn.model_selection import train_test_split
raw = load_iris()
rng = np.random.default_rng(42)
X_noisy = raw.data + rng.normal(0, 1.5, raw.data.shape)
X_train, X_test, y_train, y_test = train_test_split(
X_noisy, raw.target, test_size=0.3, random_state=42
)
model = RandomForestClassifier(n_estimators=50, random_state=42).fit(X_train, y_train)
roc = fm.roc_chart(model, X_test, y_test)
cm = fm.confusion_matrix_chart(model, X_test, y_test)
importances = fm.importance_chart(model, X_test, y_test)
report = (roc | cm) & importances
assert report.show_svg().startswith("<svg")
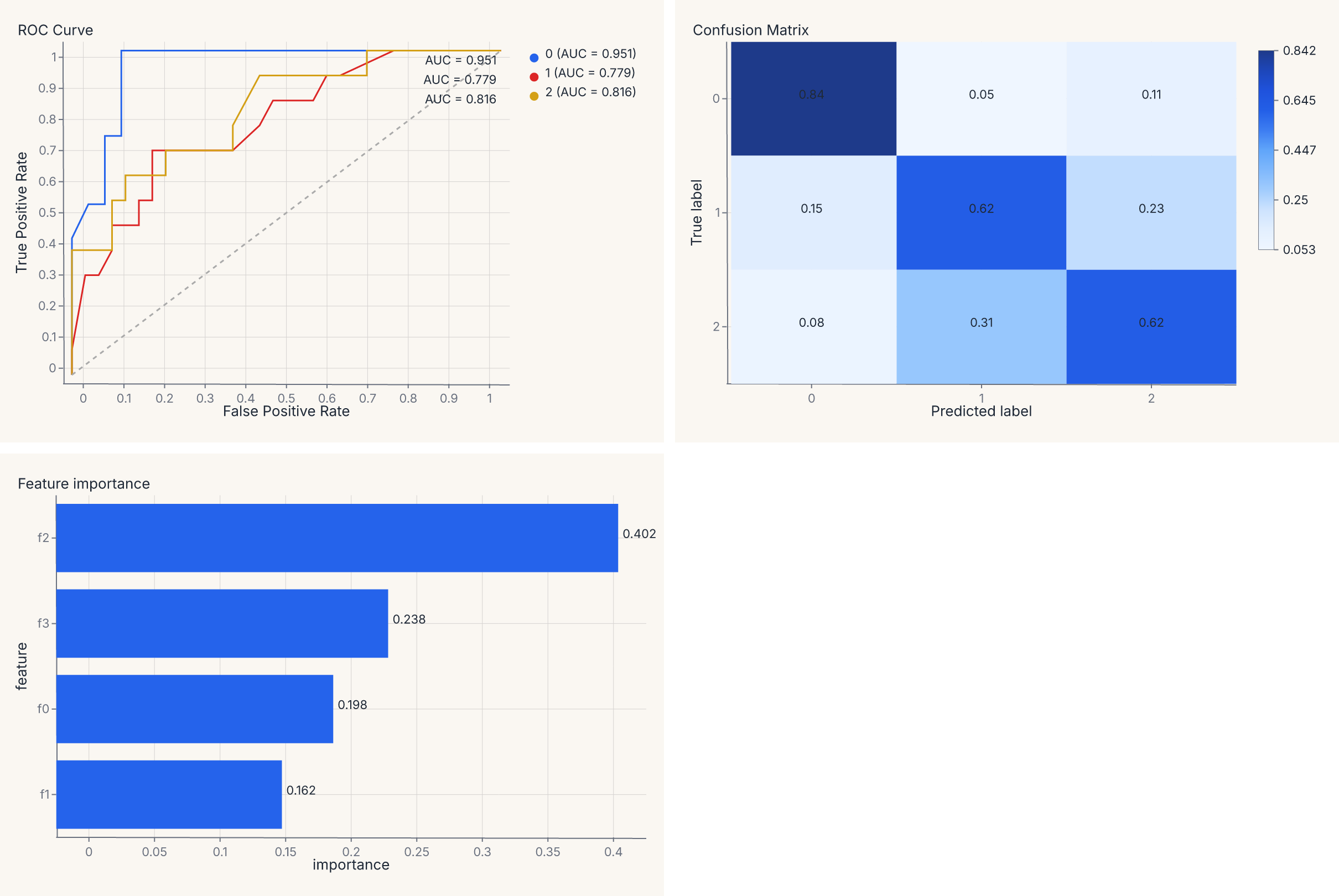
The | operator places charts side by side; & stacks vertically. The result is a single SVG with three panels — same grammar, same theme, one .save() call.
Comparing two models side by side¶
Diagnostic chart output is a regular Chart — add titles, apply themes, and compose with |:
import ferrum as fm
import numpy as np
from sklearn.datasets import load_iris
from sklearn.ensemble import RandomForestClassifier
from sklearn.linear_model import LogisticRegression
from sklearn.model_selection import train_test_split
raw = load_iris()
rng = np.random.default_rng(42)
X_noisy = raw.data + rng.normal(0, 1.5, raw.data.shape)
X_train, X_test, y_train, y_test = train_test_split(
X_noisy, raw.target, test_size=0.3, random_state=42
)
rf = RandomForestClassifier(n_estimators=50, random_state=42).fit(X_train, y_train)
lr = LogisticRegression(max_iter=500, random_state=42).fit(X_train, y_train)
roc_rf = fm.roc_chart(rf, X_test, y_test).properties(title="Random Forest")
roc_lr = fm.roc_chart(lr, X_test, y_test).properties(title="Logistic Regression")
chart = (roc_rf | roc_lr).theme(fm.themes.publication)
assert chart.show_svg().startswith("<svg")
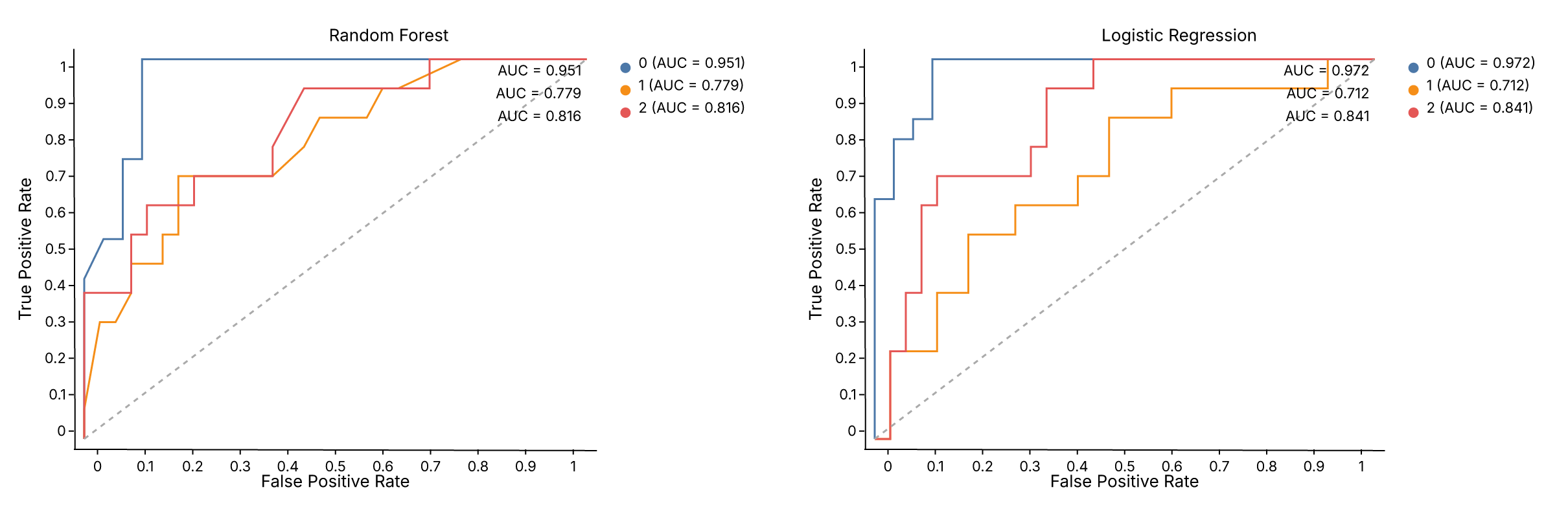
.properties(title=...) adds a title to each panel. .theme() applies to the entire composed view. The same pattern works for any diagnostic helper — swap roc_chart for calibration_chart, pr_chart, or confusion_matrix_chart.
Where to go next¶
- Marks & encodings for the full mark and encoding reference.
- Composition for layering, concatenation, and compound views.
- Figure-level helpers for one-line entry points to common patterns.
- Saving & export for output formats and rendering options.