Marks & encodings¶
Two primitives carry every Ferrum chart: marks (the geometric shapes that visualize your data) and encodings (the typed mappings from data fields to visual variables). Picking the right mark + encoding combination is most of what authoring a chart looks like.
This page is the reference for both. It covers the encoding channels, the mark families, the shorthand syntax that compresses common cases, and when to reach for what.
How a chart is assembled¶
A Ferrum chart is built by attaching a mark to a data source and declaring which columns drive which visual variables. Every chart follows the same shape:
import ferrum as fm
import polars as pl
from sklearn.datasets import load_iris
raw = load_iris()
iris = pl.DataFrame(raw.data, schema=["sepal_length", "sepal_width", "petal_length", "petal_width"]).with_columns(
species=pl.Series([raw.target_names[t] for t in raw.target])
)
chart = (
fm.Chart(iris)
.mark_point()
.encode(x="sepal_length", y="petal_length", color="species:N")
)
assert chart.show_svg().startswith("<svg")
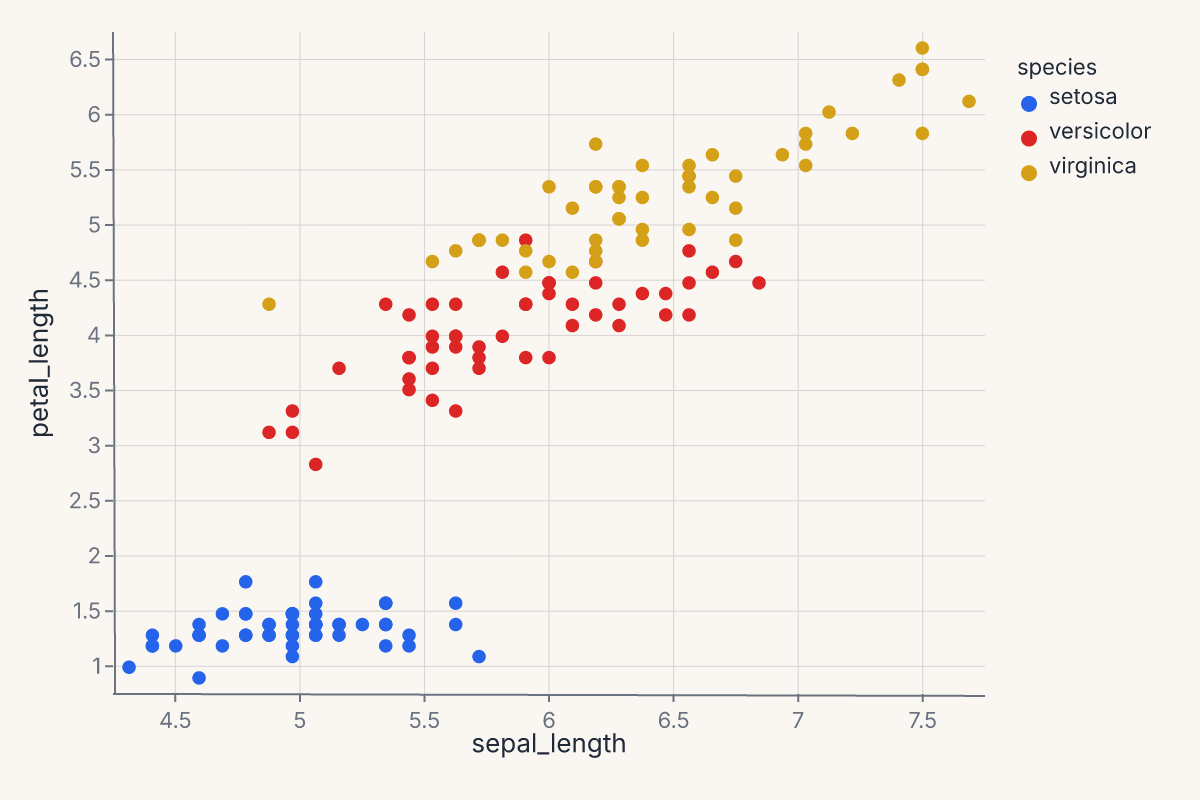
The three pieces — data, mark, encoding — compose freely. You can change the mark without touching the encoding (mark_line() instead of mark_point()), change the encoding without touching the mark, or compose multiple marks against the same encoding (see Composition).
Encoding channels¶
An encoding channel declares: this field drives this visual variable. Channels are typed by the engine: a quantitative field gets a continuous scale, a nominal field gets a categorical color palette, a temporal field gets a time scale. You can be explicit by passing an encoding object (fm.X("col", type="Q")) or use the shorthand syntax (described below).
Positional channels¶
These channels place marks in space:
| Channel | Purpose |
|---|---|
x, y |
Primary horizontal / vertical position. |
x2, y2 |
Secondary position. Used for bands, segments, intervals, error extents. |
xerror, yerror, xerror2, yerror2 |
Error extents around the primary position. |
theta, radius |
Polar coordinates. Used with CoordPolar. |
Most charts only declare x and y. The rest unlock band marks (mark_area, mark_errorband), intervals (mark_rect, mark_rule), and polar plots.
Appearance channels¶
These channels modulate how marks look:
| Channel | Purpose |
|---|---|
color |
Mark color. Continuous fields get a perceptually uniform palette; categorical fields get a discrete palette. |
fill, stroke |
Override color separately for the fill and stroke. color sets both. |
opacity, fill_opacity, stroke_opacity |
Mark opacity. |
stroke_width, stroke_dash |
Stroke styling. |
size |
Mark size. |
shape |
Mark glyph (for mark_point). |
angle |
Rotation. |
Appearance channels can take either a field name (data-driven) or a literal value (constant for all marks). Setting color="red" colors every mark red; setting color="species:N" colors marks by the species column.
Text and metadata channels¶
These channels carry information that does not directly map to position or appearance:
| Channel | Purpose |
|---|---|
text |
Text content for mark_text. |
detail |
Additional grouping that does not affect appearance — useful for keeping series separate without coloring them differently. |
tooltip, tooltip_field |
Field shown on hover. In interactive mode, renders as a tooltip overlay; in static output, becomes accessibility metadata. |
href |
URL the mark links to. |
description |
Accessibility description. |
key |
Stable identity for interactive selections. |
Faceting channels¶
These channels split the chart into small multiples:
| Channel | Purpose |
|---|---|
facet |
Single faceting variable, wrapped into a grid. |
facet_row, facet_col |
Row / column facets for a 2-D small-multiples grid. |
Faceting is structural: it produces multiple panels rather than overlaying marks. To layer marks against the same axes, use Composition.
The shorthand string syntax¶
Encodings accept a compact string syntax that handles the most common cases without explicit channel objects:
| Shorthand | Meaning |
|---|---|
"field" |
Field with inferred type (engine picks Q / N / O / T based on dtype). |
"field:Q" |
Explicitly quantitative. |
"field:N" |
Nominal (unordered categorical). |
"field:O" |
Ordinal (ordered categorical). |
"field:T" |
Temporal. |
"agg(field):Q" |
Aggregation. Examples: "mean(price):Q", "count():Q", "sum(qty):Q", "median(value):Q". |
The shorthand is purely syntactic sugar over the explicit form. fm.X("price", type="Q") and "price:Q" produce identical specs. The shorthand keeps simple cases compact; the explicit form unlocks advanced channel options.
When in doubt, use the explicit form:
import ferrum as fm
import polars as pl
from sklearn.datasets import load_iris
raw = load_iris()
iris = pl.DataFrame(raw.data, schema=["sepal_length", "sepal_width", "petal_length", "petal_width"]).with_columns(
species=pl.Series([raw.target_names[t] for t in raw.target])
)
chart = (
fm.Chart(iris)
.mark_point()
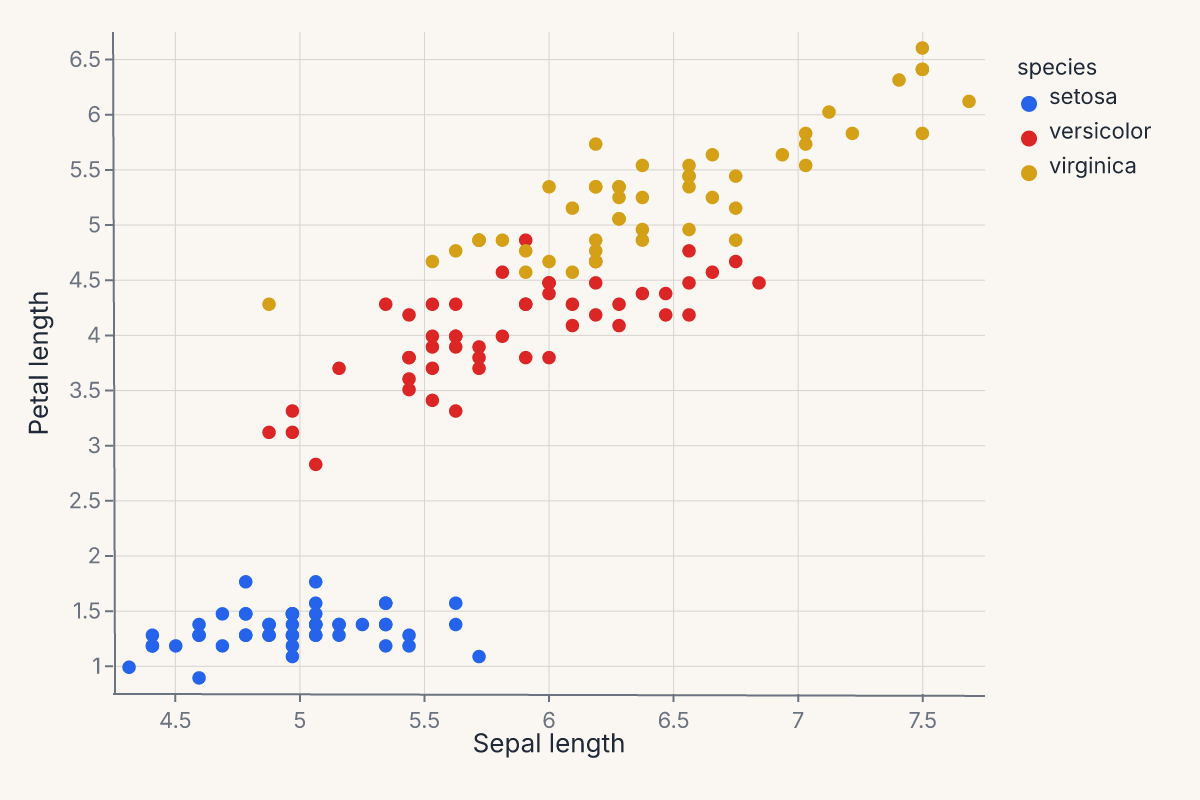
.encode(
x=fm.X("sepal_length", type="Q", title="Sepal length"),
y=fm.Y("petal_length", type="Q", title="Petal length"),
color=fm.Color("species", type="N", title="Species"),
)
)
assert chart.show_svg().startswith("<svg")
Mark families¶
Ferrum ships 54 mark methods on Chart. They group into families by what they're for.
Primitive marks¶
The geometric building blocks. Use these when you want direct control over what gets drawn.
| Method | Geometry |
|---|---|
mark_point() |
Discrete points. The default scatter mark. |
mark_line() |
Polyline connecting points in order. |
mark_bar() |
Vertical or horizontal bars. |
mark_area() |
Filled area, optionally banded with y2. |
mark_rule() |
Reference lines (often horizontal or vertical). |
mark_tick() |
Short ticks, often used for rug plots. |
mark_rect() |
Rectangular cells. Used for heatmaps and intervals. |
mark_text() |
Text labels (paired with the text encoding). |
mark_label() |
Positioned text labels with collision avoidance (avoid_overlap=True). |
mark_image() |
Image tiles from URL fields. |
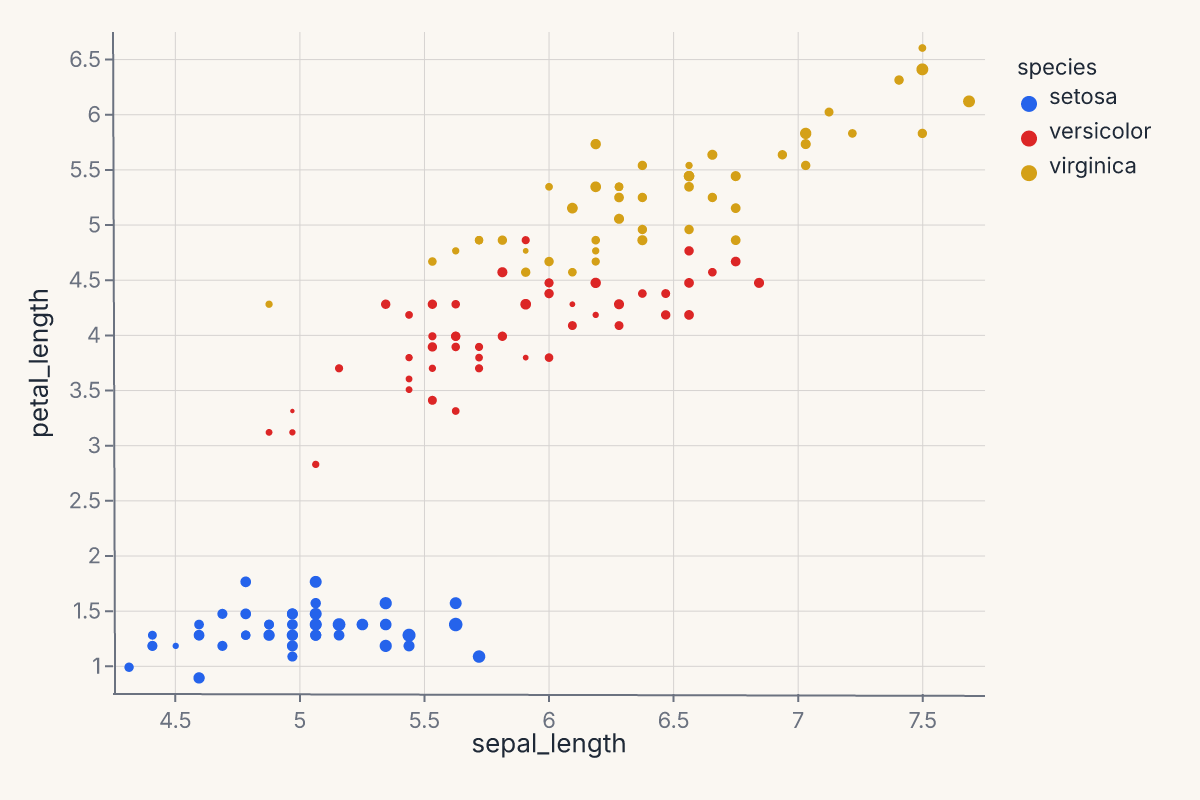
Example — basic scatter:
import ferrum as fm
import polars as pl
from sklearn.datasets import load_iris
raw = load_iris()
iris = pl.DataFrame(raw.data, schema=["sepal_length", "sepal_width", "petal_length", "petal_width"]).with_columns(
species=pl.Series([raw.target_names[t] for t in raw.target])
)
chart = (
fm.Chart(iris)
.mark_point()
.encode(
x="sepal_length",
y="petal_length",
color="species:N",
size="sepal_width",
)
)
assert chart.show_svg().startswith("<svg")
Statistical marks¶
These marks compute a transform on your data before rendering — KDE, binning, smoothing, contours, quantile-quantile reference, or arbitrary functions. The transform happens in Rust, declared in the chart spec.
| Method | Transform |
|---|---|
mark_density() |
1-D kernel density estimate. |
mark_histogram() |
Binned counts or densities. |
mark_smooth() |
LOESS / GLM / logistic regression overlay. |
mark_contour() |
2-D density contours. |
mark_qq() |
Quantile-quantile plot against a reference distribution. |
mark_violin() |
Symmetric KDE per group. |
mark_function() |
Plot an arbitrary f(x) over a domain. |
Example — 1-D kernel density estimate:
import ferrum as fm
import polars as pl
from sklearn.datasets import load_iris
raw = load_iris()
iris = pl.DataFrame(raw.data, schema=["sepal_length", "sepal_width", "petal_length", "petal_width"]).with_columns(
species=pl.Series([raw.target_names[t] for t in raw.target])
)
chart = (
fm.Chart(iris)
.mark_density(bandwidth="scott")
.encode(x="sepal_length")
)
assert chart.show_svg().startswith("<svg")
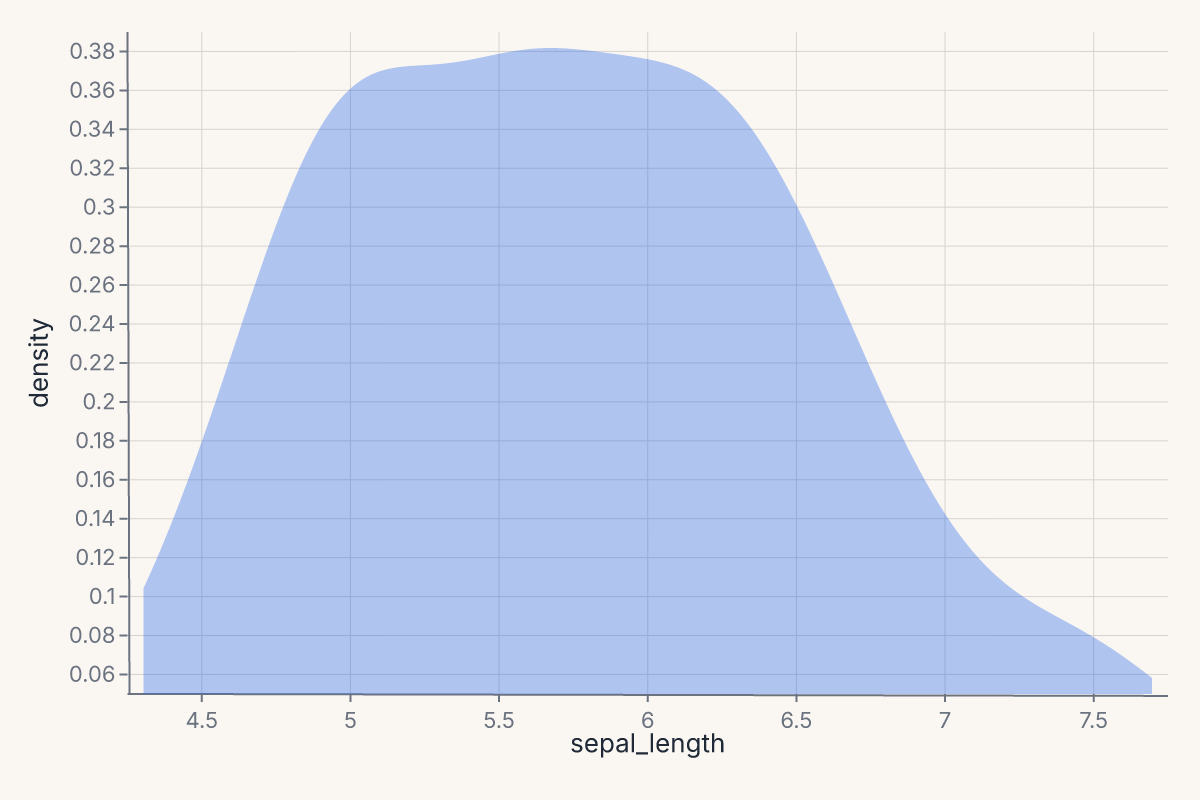
Stat marks are described in detail in Stats in the rendering pipeline.
Grouped transforms with groupby¶
Statistical marks compute their transform over the entire dataset by default. To compute independently per group — one LOESS line per species, one KDE per category — pass groupby=:
import ferrum as fm
import polars as pl
from sklearn.datasets import load_iris
raw = load_iris()
iris = pl.DataFrame(raw.data, schema=["sepal_length", "sepal_width", "petal_length", "petal_width"]).with_columns(
species=pl.Series([raw.target_names[t] for t in raw.target])
)
chart = (
fm.Chart(iris)
.mark_smooth(method="loess", groupby="species")
.encode(x="sepal_length", y="petal_length", color="species:N")
)
assert chart.show_svg().startswith("<svg")
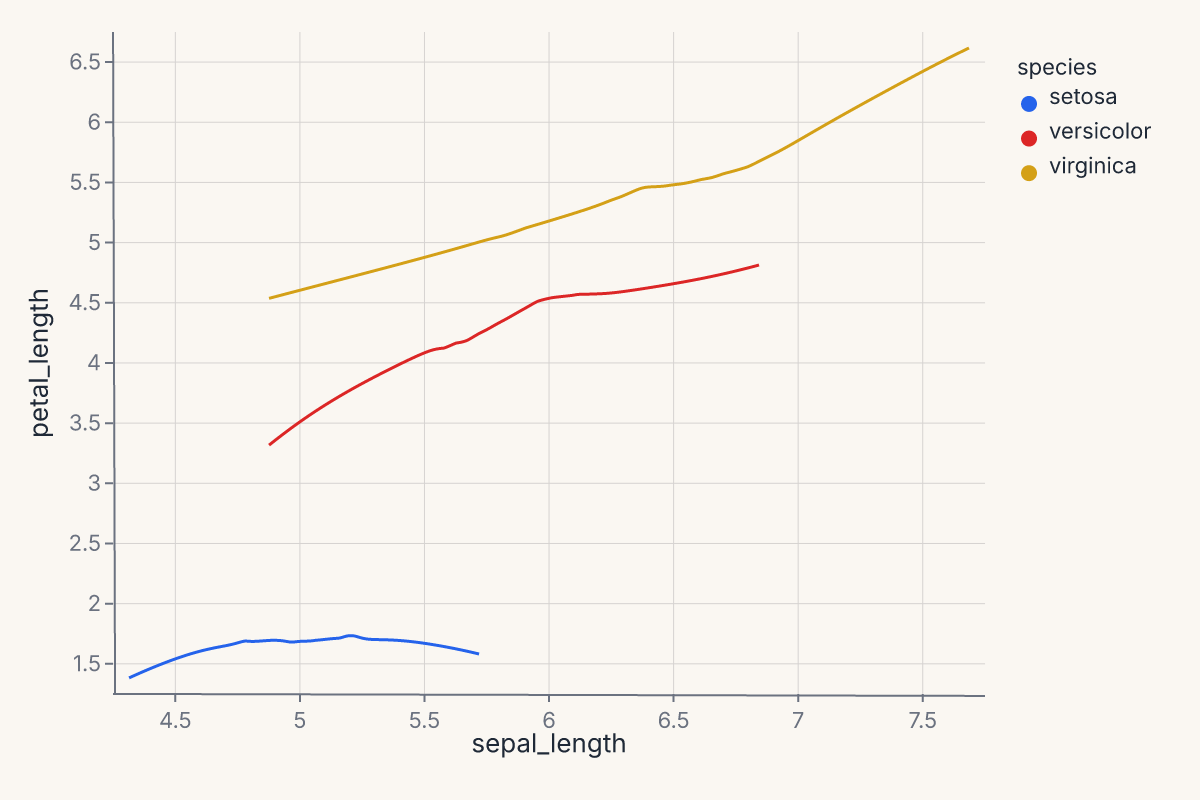
The groupby parameter is available on mark_smooth, mark_density, and mark_histogram. The group column is preserved in the transform output so downstream color= encoding maps each group to a distinct visual.
This is especially important when layering a statistical mark with a scatter via + — without groupby, the transform runs over all data combined and produces a single aggregate line.
Distribution-summary marks¶
For comparing categorical distributions at a glance:
| Method | Geometry |
|---|---|
mark_boxplot() |
Tukey boxplot — quartiles, whiskers, outliers. |
mark_boxen() |
Letter-value boxplot — more quantiles for larger samples. |
mark_swarm() |
Beeswarm jitter (categorical scatter without overlap). |
Example — boxplot by species:
import ferrum as fm
import polars as pl
from sklearn.datasets import load_iris
raw = load_iris()
iris = pl.DataFrame(raw.data, schema=["sepal_length", "sepal_width", "petal_length", "petal_width"]).with_columns(
species=pl.Series([raw.target_names[t] for t in raw.target])
)
chart = (
fm.Chart(iris)
.mark_boxplot()
.encode(x="species:N", y="sepal_length")
)
assert chart.show_svg().startswith("<svg")
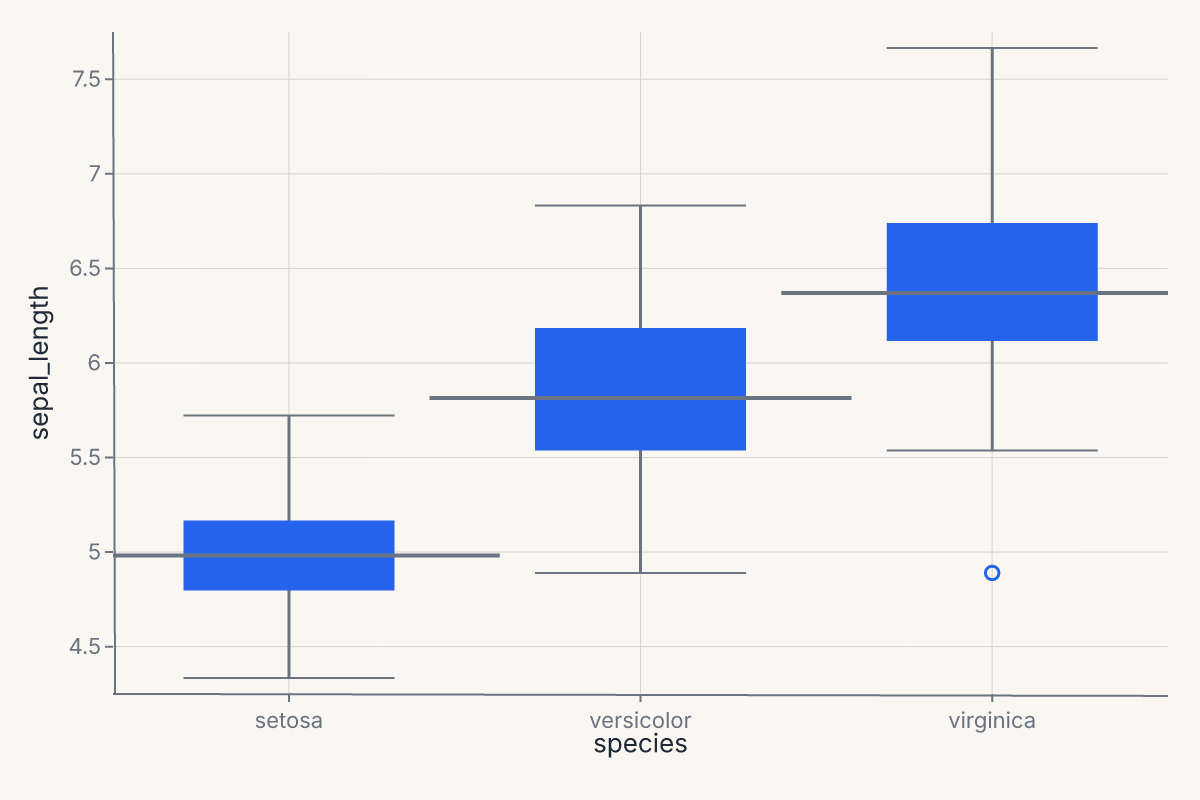
Uncertainty marks¶
For showing the spread around an estimate:
| Method | Geometry |
|---|---|
mark_errorbar() |
Error bars with optional terminal ticks. |
mark_errorband() |
Filled band between y and y2. |
mark_ribbon() |
Continuous band, typically over a line. |
Scale-aware marks¶
For large data where vector marks would overwhelm the renderer:
| Method | Geometry |
|---|---|
mark_raster() |
Pre-aggregated rectangular grid. |
mark_hex() |
Hexagonal binning. |
Both fall back to explicit raster output regardless of the renderer in use. See Performance & scale for when to reach for them.
Model-diagnostic marks¶
Phase 10 introduced a family of model-evaluation marks that work with ModelSource to produce ROC curves, calibration plots, residuals, SHAP summaries, and similar diagnostics — see the Model diagnostics page. These marks live alongside the rest because, structurally, a ROC curve is a chart. The full list (selected): mark_residuals, mark_prediction_error, mark_roc, mark_pr, mark_calibration, mark_gain, mark_lift, mark_confusion, mark_importance, mark_shap_beeswarm, mark_shap_bar, mark_pdp, mark_learning_curve.
For most diagnostic use cases you should reach for the figure-level helpers (roc_chart, calibration_chart, etc.) covered in Figure-level helpers rather than calling the marks directly. The marks exist for cases where you want to drop into the grammar and compose custom diagnostic views.
Picking a mark¶
A quick decision guide for the common cases:
- One variable, looking at distribution shape?
mark_density()ormark_histogram(). - One variable across groups?
mark_boxplot()(ormark_violin()for symmetric KDEs,mark_swarm()for full points). - Two variables, looking at relationship?
mark_point()(low cardinality),mark_hex()(high cardinality),mark_smooth()overlaid on points (with a regression line). - Two variables over time?
mark_line(), optionally withmark_ribbon()ormark_errorband()for uncertainty. - Counts by category?
mark_bar(). Addencode(color="...")and a stacking position adjustment for stacked bars. - A discrete grid of values?
mark_rect(). The same primitive serves heatmaps and binned 2-D histograms. - Model diagnostic? Use the figure-level helpers in Figure-level helpers and Model diagnostics rather than calling diagnostic marks directly.
A complete example¶
Combining multiple marks against one data source with shared encodings (full composition is covered on the Composition page):
import ferrum as fm
import polars as pl
from sklearn.datasets import load_iris
raw = load_iris()
iris = pl.DataFrame(raw.data, schema=["sepal_length", "sepal_width", "petal_length", "petal_width"]).with_columns(
species=pl.Series([raw.target_names[t] for t in raw.target])
)
points = (
fm.Chart(iris)
.mark_point(opacity=0.6)
.encode(x="sepal_length", y="petal_length", color="species:N")
)
trend = (
fm.Chart(iris)
.mark_smooth(method="loess")
.encode(x="sepal_length", y="petal_length", color="species:N")
)
combined = points + trend
assert combined.show_svg().startswith("<svg")
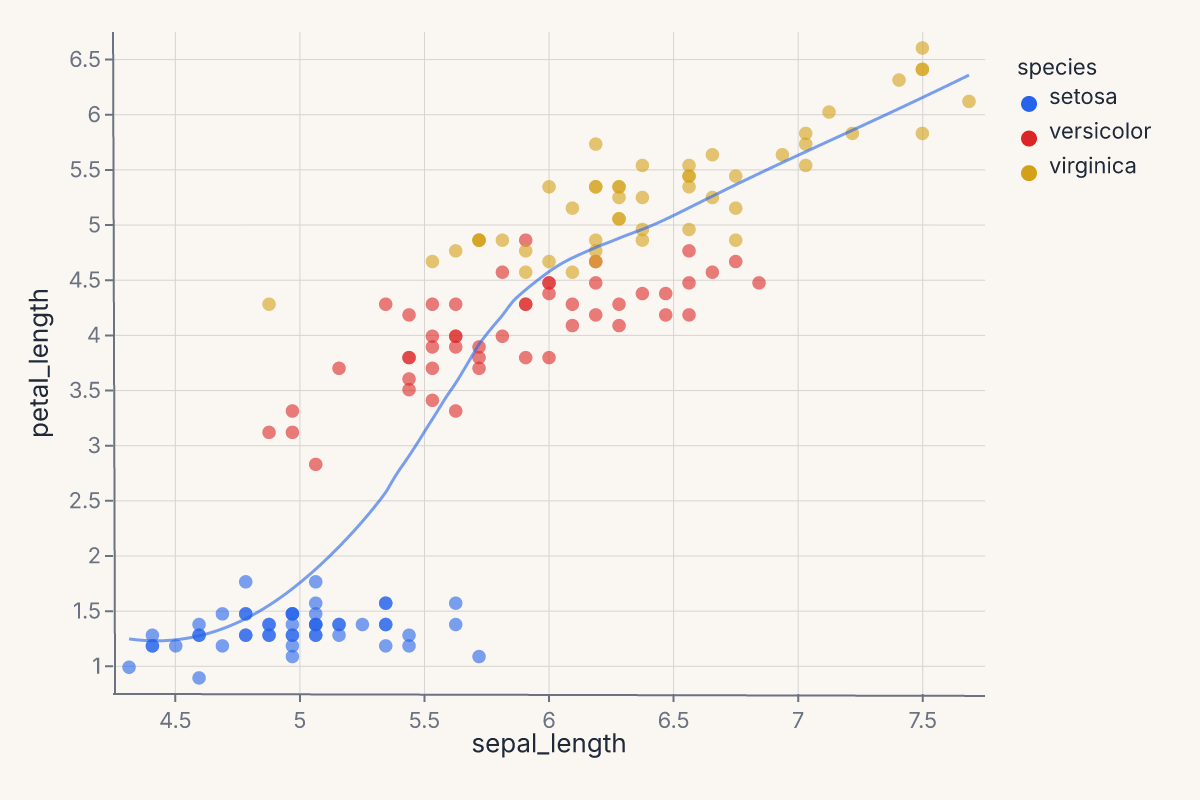
This puts a per-species LOESS overlay on top of a scatter. Same encoding, two marks, one layered chart — the + operator on Chart produces a layered view that renders both marks against the same axes.
Where to go next¶
- Composition for how to combine multiple marks and charts into compound views (
Layer,HConcat,VConcat,JointChart, etc.). - Themes for changing how marks look without changing the chart spec.
- Figure-level helpers for one-line entry points to common chart patterns.
- Stats in the rendering pipeline for the design rationale behind statistical marks.
- The API Reference for the full method signatures of every mark and encoding channel.