First plot¶
This page gets you from zero to a rendered chart in under a minute. By the end you'll have a scatter plot, a layered chart, and a saved SVG — and you'll know the three-piece pattern that every Ferrum chart follows.
The pattern¶
Every Ferrum chart is data + mark + encoding:
import ferrum as fm
import polars as pl
from sklearn.datasets import load_iris
raw = load_iris()
iris = pl.DataFrame(raw.data, schema=["sepal_length", "sepal_width", "petal_length", "petal_width"]).with_columns(
species=pl.Series([raw.target_names[t] for t in raw.target])
)
chart = (
fm.Chart(iris)
.mark_point()
.encode(x="sepal_length", y="petal_length", color="species:N")
)
assert chart.show_svg().startswith("<svg")
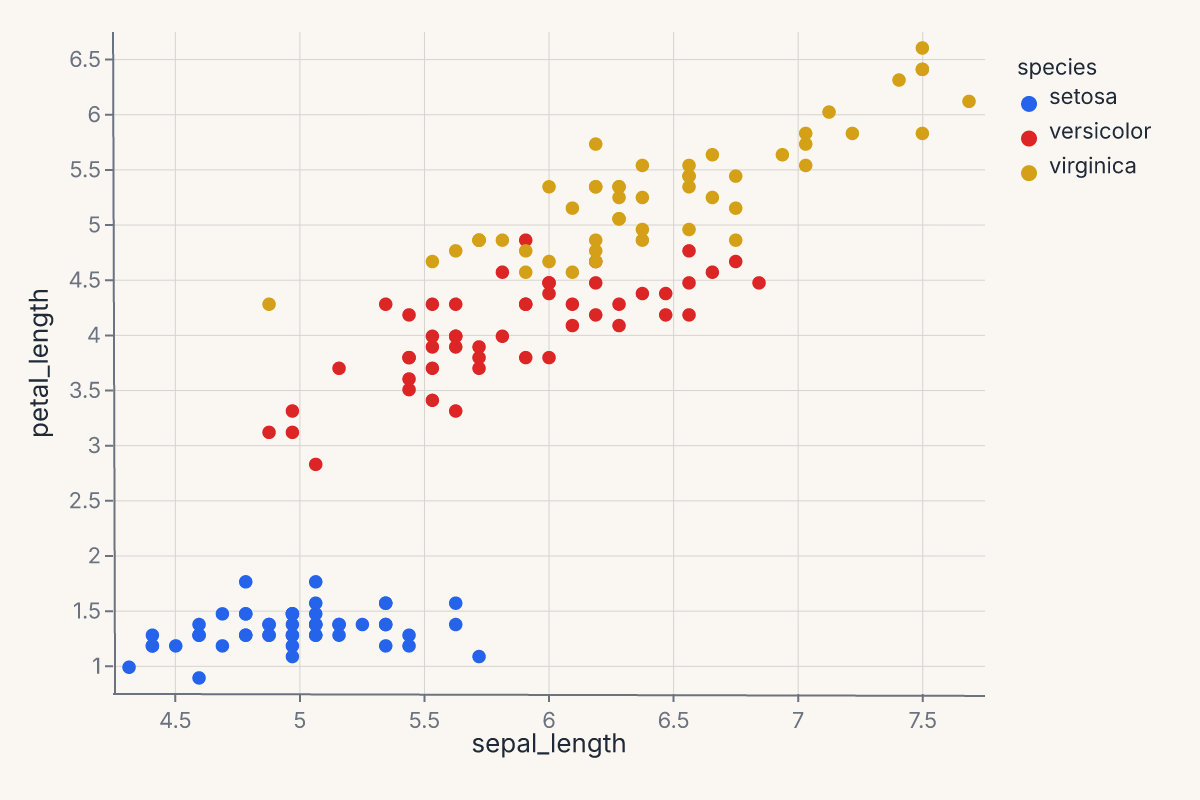
That's the whole thing: Chart(data) binds your DataFrame, .mark_point() picks the geometry, .encode(...) maps columns to visual channels. The result is a Chart object — call .show_svg() to render it, .save() to write it to disk, or just display it in a Jupyter notebook (where it renders automatically).
Add a trend line¶
Want a regression overlay? Layer it with +:
import ferrum as fm
import polars as pl
from sklearn.datasets import load_iris
raw = load_iris()
iris = pl.DataFrame(raw.data, schema=["sepal_length", "sepal_width", "petal_length", "petal_width"]).with_columns(
species=pl.Series([raw.target_names[t] for t in raw.target])
)
points = (
fm.Chart(iris)
.mark_point(opacity=0.6)
.encode(x="sepal_length", y="petal_length", color="species:N")
)
trend = (
fm.Chart(iris)
.mark_smooth(method="loess", groupby="species")
.encode(x="sepal_length", y="petal_length", color="species:N")
)
chart = points + trend
assert chart.show_svg().startswith("<svg")
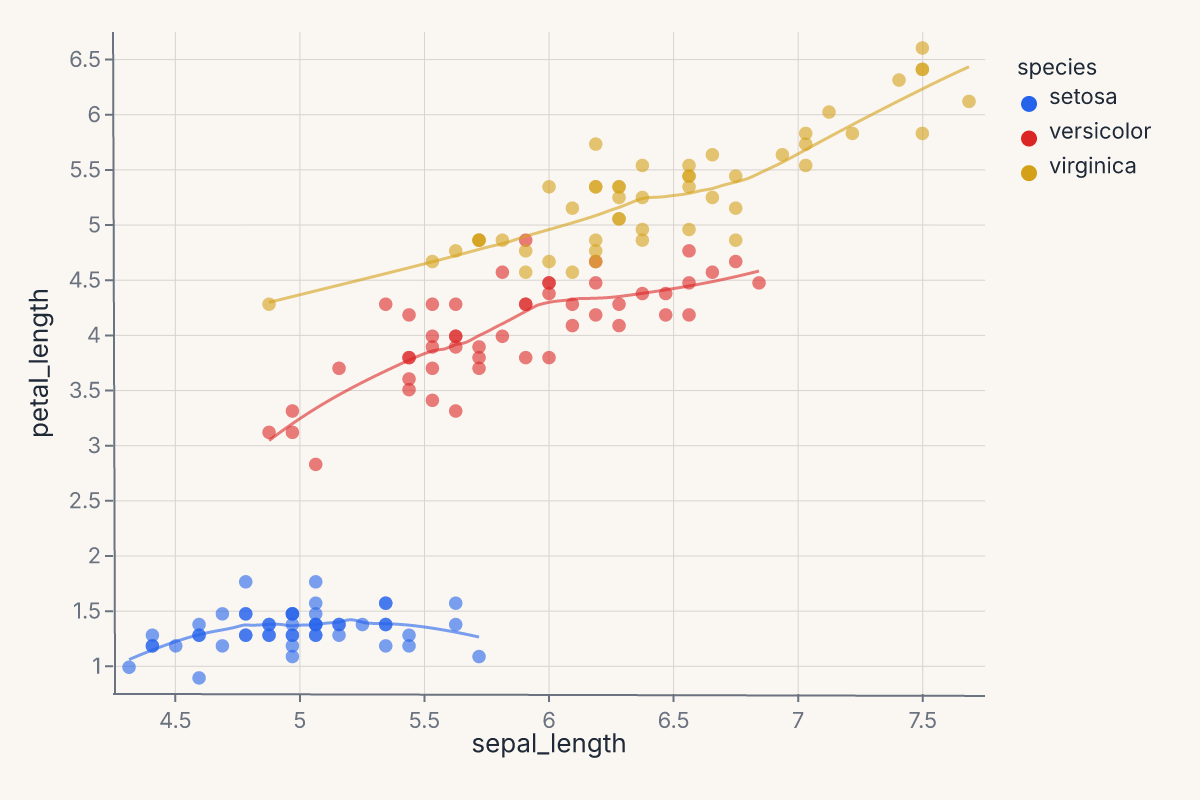
The + operator always layers — both marks share the same axes. The LOESS smooth is computed in Rust; you declared what you wanted, not how to compute it.
Try a different mark¶
Different questions call for different marks. The pattern is always the same — data, mark, encoding:
import ferrum as fm
import polars as pl
from sklearn.datasets import load_iris
raw = load_iris()
iris = pl.DataFrame(raw.data, schema=["sepal_length", "sepal_width", "petal_length", "petal_width"]).with_columns(
species=pl.Series([raw.target_names[t] for t in raw.target])
)
chart = (
fm.Chart(iris)
.mark_boxplot()
.encode(x="species:N", y="sepal_length", color="species:N")
)
assert chart.show_svg().startswith("<svg")
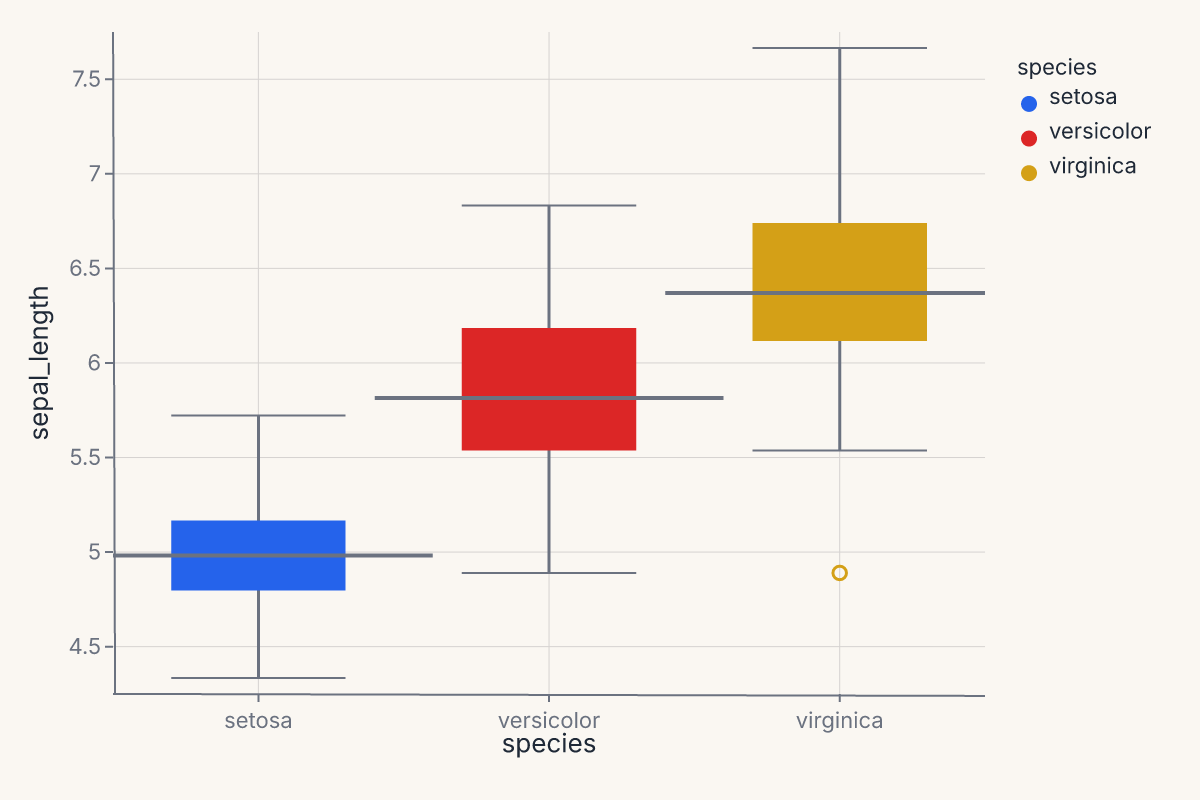
Apply a theme¶
Themes are one method call:
import ferrum as fm
import polars as pl
from sklearn.datasets import load_iris
raw = load_iris()
iris = pl.DataFrame(raw.data, schema=["sepal_length", "sepal_width", "petal_length", "petal_width"]).with_columns(
species=pl.Series([raw.target_names[t] for t in raw.target])
)
chart = (
fm.Chart(iris)
.mark_point()
.encode(x="sepal_length", y="petal_length", color="species:N")
.theme(fm.themes.publication)
)
assert chart.show_svg().startswith("<svg")
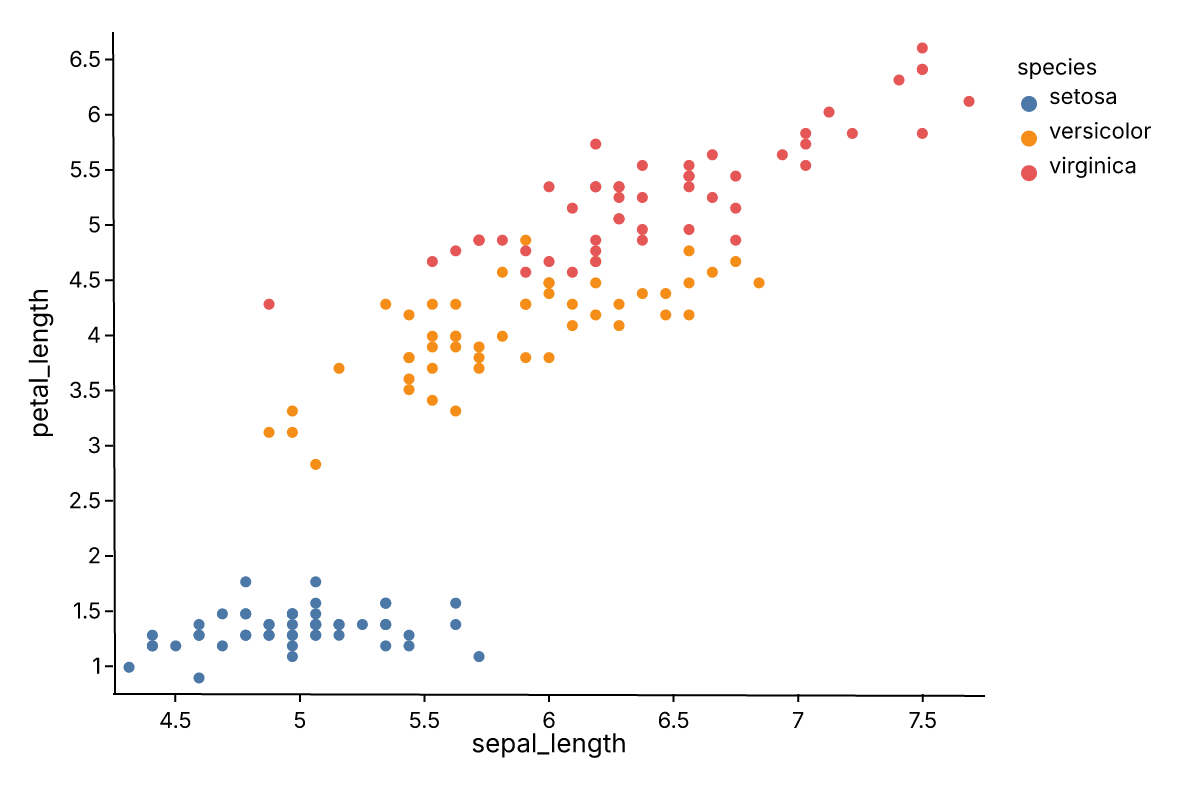
Ferrum ships twelve built-in themes — from Paper Ink (the warm default) to dark, publication, and editorial styles.
What just happened¶
In four snippets you used:
- Data binding —
fm.Chart(iris)accepts polars, pandas, modin, cuDF, dask, ibis, pyarrow, or dict-of-arrays. One constructor. - Marks —
mark_point(),mark_smooth(),mark_boxplot(). Ferrum has 28+ marks covering primitives, statistical transforms, distributions, and model diagnostics. - Encodings —
x,y,color. Shorthand strings like"species:N"set the type (Nominal); the full formfm.X("field", type="Q", title="...")gives finer control. - Composition —
+layers marks on shared axes.|and&concatenate charts side-by-side or stacked. - Themes —
.theme(fm.themes.publication)swaps the entire visual style without touching the data or encoding.
Where to go next¶
- Marks & encodings — the full mark and encoding reference.
- Composition — layering, concatenation, joint charts, repeat grids.
- Themes — the twelve built-in themes, custom themes, and scoped defaults.
- Figure-level helpers — one-line entry points for common chart patterns.
- Model diagnostics — ROC curves, confusion matrices, SHAP — all as charts.